Hi, what are the best practices in reading/working with large geospatial data (netcdf, hdf5, etc.)?
There is one recommendation that is often mentioned, which is to pay attention to the index order.
Are there other suggestions?
What R and Python packages are most optimal to work with these (very large) geospatial data files in a cluster environment?
Are their any publicly accessible tutorials on working with very large geospatial data?
Here is a tutorial I found that I think is good:
And I think Pangeo is a good place to find out what is community ‘best practices’.
https://pangeo.io
From my own experience in profiling user code, (this is just for Python), the thing that makes things super horribly slow is not using numpy properly. To get the speed of numpy, you need to make sure everything is a numpy object. For example, one researcher was reading in satellite data in netCDF form and it created an xarray object. xarray objects have metadata and…well, they aren’t numpy arrays. It make loops soooo slow, and he had a double loop. I did something like
my_data = xr.DataArray.to_numpy(data)
and got 33x speed up.
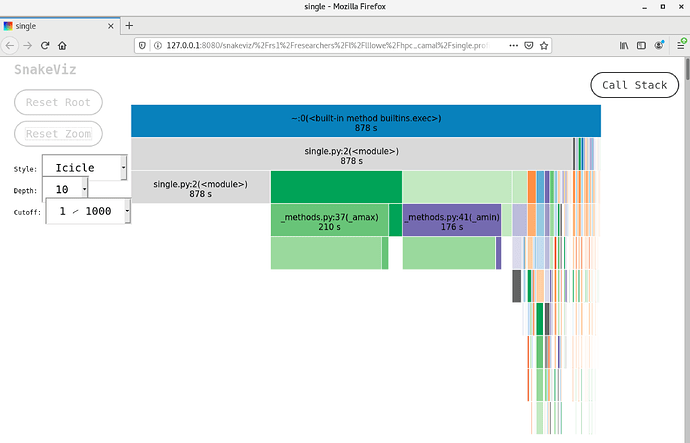
Similarly, for one user, they thought they were using numpy array, and they were using numpy library, but somehow the variable didn’t register as numpy and max/min functions were taking forever. I found the problem by using the ‘snakeviz’ profiler. Please look up and confirm the current proper syntax, but from what notes I took, you do something like this:
For a Python script named mycode.py, do a profile and call the output file mycode.prof:
python -m cProfile -o mycode.prof mycode.py
Then use the command snakeviz
snakeviz mycode.prof
and it will open a webbrowser that has the profiling information.
For the second case I mentioned, all the time was spent in the ‘min/max’ function. The array was not officially ‘numpy’ format. Before fixing it, it took over six minutes, and afterwards, only a few seconds. So just check the data types. I found the first problem just by checking the data type, and saw it had metadata attached.
I’d suggest looking into Python’s multiprocessing library. When working with computationally intensive problems in a high-performance environment, it’s a good idea to benchmark your script. Then try throwing more computing resources at it and see if you get quicker results that way. Try taking advantage of parallel computing (using the multiprocessing library, for example) and see if you can get some significant speed-ups.
Here’s a quick guide to multiprocessing in Python: https://www.sitepoint.com/python-multiprocessing-parallel-programming/
And Penn State has some intro-level info in an open textbook regarding parallel processing and geospatial data that might be helpful: https://www.e-education.psu.edu/geog489/node/2253
This is very useful! Thank you, --Katia
Thank you, This is awesome! --Katia
Hi, apart from paying attention to the index order, other best practices to consider when working with geospatial data include 1. Using appropriate data structures. For large datasets, it is best to use data structures that are designed for efficient storage and access of large amounts of data. and 2. Using appropriate tools.
Thank you @mogunleye . Any specific recommendations about “appropriate tools” and “appropriate data structures”? Or links that describe these tools and structures?
Thank you,
–Katia